- ���(y��) ���Մ�(d��ng)�B(t��i) �˲��Ј�(ch��ng) �¼��g(sh��)���� �Ї�(gu��)�ƌW(xu��)�� ��չ�_(t��i) ���v��ֱ�� ��(hu��)չ���� �r(ji��)���� ���g(sh��)��Ӎ ���M(f��i)ԇ��

-

����ͨ��
����ץס�����Ƽ�
����(d��ng)���}��
Arraystar rG4оƬ�����iRNA��(d��ng)�B(t��i)�Y(ji��)��(g��u)�W��
�����w�� �� �� С �� �r(sh��)�g��2025��12��16�� ��(l��i)Դ����������
�����]��
����Arraystar rG4оƬ���g(sh��)���H�܉���Ч���@�͙z�y(c��)�(x��)���е�rG4��ͬ�r(sh��)Ҳ������DMS�T��(d��o)��������a(ch��n)����ƫ��Ķ��@�������rG4 �����D�V�����Y(ji��)���Ĝ�(zh��n)�_�ԺͿɿ��ԡ�
һ����(ji��n)��
RNA G-��(li��n)�w (RNA G-quadruplexes��rG4)���ɸ����B(ni��o)���� (G) ��RNA����ͨ�^(gu��)Hoogsteen���I�γɵ�һ�N�ǽ�(j��ng)�����(j��)�Y(ji��)��(g��u)��ԓ�Y(ji��)��(g��u)�ɶѯB��G-�ķ��wƽ�昋(g��u)�ɣ�����K⁺�Ȇr(ji��)�(y��ng)�x�ӷ�(w��n)�����D1����rG4�Ą�(d��ng)�B(t��i)�Y(ji��)��(g��u)�D(zhu��n)׃���{(di��o)��RNA�D(zhu��n)�[1]��Ⱦɫ�|(zh��)�������ļ��[2]��miRNAǰ�w�ӹ�[3]��mRNA���g[4, 5]�Լ� mRNA ��(w��n)����[6]�����⣬rG4 ߀���cm7G [3]��o8G[7]��m6A[8, 9]�� RNA ���ͬ�{(di��o)�ػ�����_(d��)��rG4�γɰl(f��)��ʧ�{(di��o)���t��(hu��)Ӱ푑�(y��ng)������(y��ng) [7]�����Y������_(d��)�{(di��o)�� [10, 11]�����c����ɭ����·���w�V������ϵ�y(t��ng)ή�s�аl(f��)���Ħ�-ͻ�|�˵��ۼ����P(gu��n)[12]��
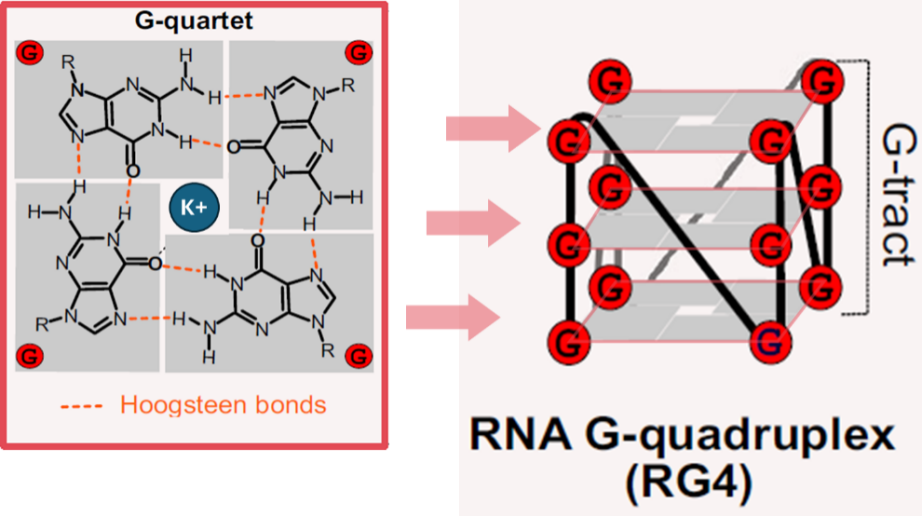
�D1. �ڸ����B(ni��o)���ʵ�RNA�����У��Ă�(g��)�B(ni��o)����ͨ�^(gu��)Hoogsteen�I�Y(ji��)���γ�G-�ķ��wƽ�棬G-�ķ��wƽ��ѯB�γ�RNA G-��(li��n)�w��RG4��[13]��
Arraystar rG4оƬ���g(sh��)�ɾ���(zh��n)�����D(zhu��n)䛽M�е�rG4 �Y(ji��)��(g��u)�����е��P(gu��n)�I���E�������w��(n��i)��������� (Dimethyl sulfate, DMS) ̎�����w��RNA���ۯB���Լ�ʹ�ÿ�G4���w��BG4���M(j��n)���H���Բ��@���S�����@���ĺ��� rG4�Y(ji��)��(g��u)�� RNA�M(j��n)��ȥ����̎������ȥ�� DMS ̎�����a(ch��n)���ĸ��a(ch��n)��m1A/m3C����������ɵęz�y(c��)�ɔ_��������ø��`���ȵ�Arraystar rG4 оƬ̽ᘌ�(du��)rG4-RNA�M(j��n)�ж���������Arraystar rG4оƬ���g(sh��)���H�܉���Ч���@�͙z�y(c��)�(x��)���е�rG4��ͬ�r(sh��)Ҳ������DMS�T��(d��o)��������a(ch��n)����ƫ��Ķ��@�������rG4 �����D�V�����Y(ji��)���Ĝ�(zh��n)�_�ԺͿɿ��ԡ�

�������g(sh��)��(y��u)��(sh��)
�w��(n��i)DMS̎���c�w�����ۯB��(f��)�F(xi��n)���挍(sh��)��rG4�Y(ji��)��(g��u)
ʹ�ø��H���Ե�BG4���w�خ��Ը�������rG4�Y(ji��)��(g��u)��RNA
Demthylation̎��ȥ����DMS ̎���ĸ��a(ch��n)��m1A/m3C����ɵęz�y(c��)�ɔ_
Arraystar оƬ�����`���ęz�y(c��)rG4 RNA, ����RNA�y(c��)��o(w��)����(zh��n)�_�z�y(c��)�ĵ��S��RNA
������(sh��)�(y��n)����
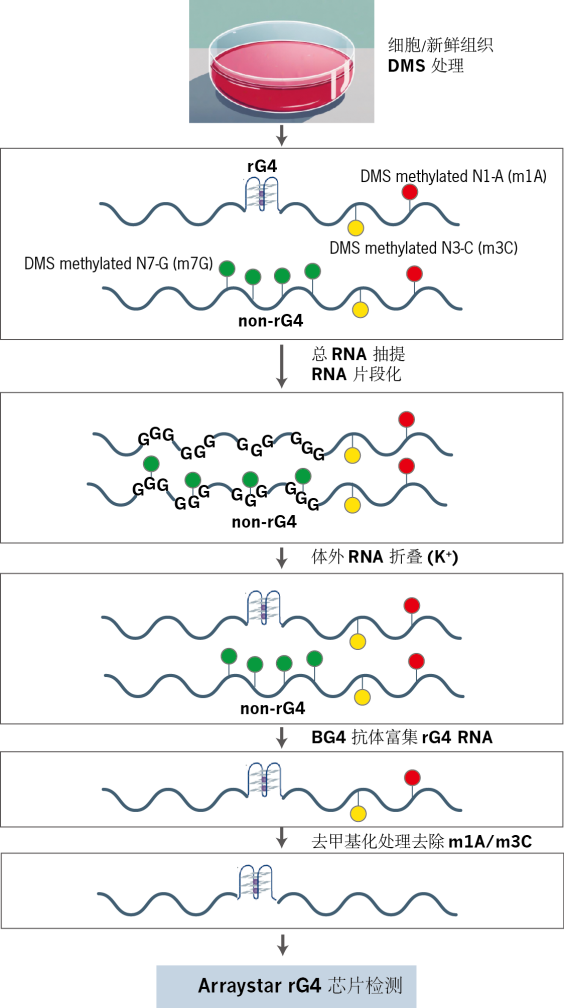
�D2. rG4оƬ��(sh��)�(y��n)���̈D��
1. �w��(n��i) DMS ̎��
���B(y��ng)��(x��)����(j��ng)�����������DMS��̎������(du��) RNA �е������ʣ�A��������ण�C������ rG4�ۯB�^(q��)����B(ni��o)���ʣ�G���M(j��n)�м������rG4 �Y(ji��)��(g��u)��(n��i)�� G �A������gλ�費��Ӱ푣��Ķ�������δ������B(t��i)��
2. RNA �����c�w�����ۯB
���x�� RNA ���M(j��n)��Ƭ�λ�̎�����S���ں�K⁺�ľ��_�wϵ�н�(j��ng)�v׃��-��(f��)���^(gu��)�̡��˲��E�H���S��(x��)����(n��i)ԭ�е� rG4 �^(q��)���خ������ۯB���������^(q��)�����ڔy��m7G����o(w��)���γɷ�(w��n)���Y(ji��)��(g��u)��
3. rG4 ���߳���
���ÿ� G-��(li��n)�w���w��BG4����(du��)�� rG4 �Y(ji��)��(g��u)�� RNA �M(j��n)�����߳�����IP������(sh��)�F(xi��n)Ŀ��(bi��o)���ӵ��خ��Ը�����
4. ȥ�����c���D(zhu��n)�
������ RNA ��(j��ng)ȥ����ø̎������� DMS �T��(d��o)�ĸ��a(ch��n)��� m¹A/m³C���������z�y(c��)ƫ��S��ͨ�^(gu��)���D(zhu��n)䛺ϳ��p� cDNA��������T7 ����(d��ng)�����С�
5. �ɹ��(bi��o)ӛ cRNA �ϳ�
�� cDNA ��ģ�壬T7 RNA �ۺ�ø���w���D(zhu��n)䛷���(y��ng)������Cy3-CTP �ɹ�Ⱦ�ϣ����Ɏ�Cy3 ��(bi��o)ӛ�ķ��x cRNA��
6. оƬ�s���c��(sh��)��(j��)����
��(bi��o)ӛ�� cRNA �cArraystar rG4 оƬ�s����ͨ�^(gu��)�ɹ���̖(h��o)���������D(zhu��n)䛽M��rG4 �Y(ji��)��(g��u)�ķֲ��c�S�ȡ�
�ġ�оƬ����(sh��)
Arraystar ��� rG4 оƬ����(sh��)
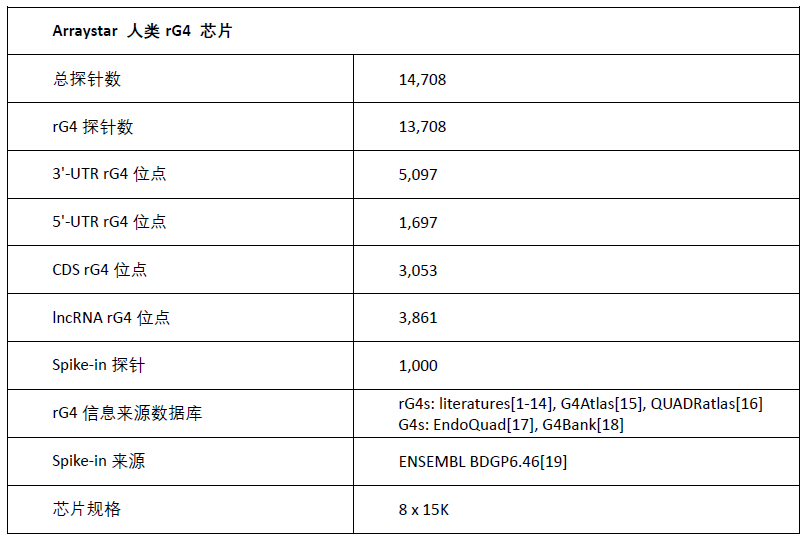
RNA G-quadruplex (rG4) ��(sh��)��(j��)��(k��)
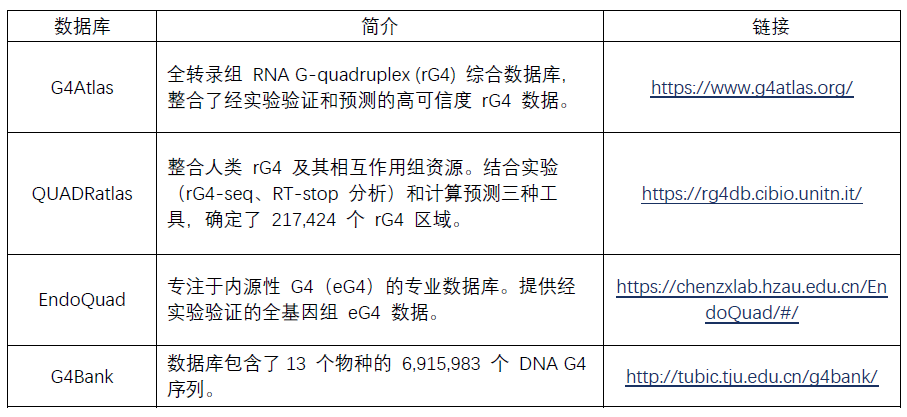
�����īI(xi��n)
1. Yari H et al: LncRNA REG1CP promotes tumorigenesis through an enhancer complex to recruit FANCJ helicase for REG3A transcription. Nat Commun 2019, 10(1):5334.[PMID: 31767869]
2. Lee YW, Weissbein U, Blum R, Lee JT: G-quadruplex folding in Xist RNA antagonizes PRC2 activity for stepwise regulation of X chromosome inactivation. Mol Cell 2024, 84(10):1870-1885 e1879.[PMID: 38759625]
3. Pandolfini L et al: METTL1 Promotes let-7 MicroRNA Processing via m7G Methylation. Mol Cell 2019, 74(6):1278-1290 e1279.[PMID: 31031083]
4. Arora A, Suess B: An RNA G-quadruplex in the 3' UTR of the proto-oncogene PIM1 represses translation. RNA Biol 2011, 8(5):802-805.[PMID: 21734463]
5. Song J, Perreault JP, Topisirovic I, Richard S: RNA G-quadruplexes and their potential regulatory roles in translation. Translation (Austin) 2016, 4(2):e1244031.[PMID: 28090421]
6. Rouleau S et al: 3' UTR G-quadruplexes regulate miRNA binding. RNA 2017, 23(8):1172-1179.[PMID: 28473452]
7. Ma Y et al: RNA G-Quadruplex within the 5'-UTR of FEN1 Regulates mRNA Stability under Oxidative Stress. Antioxidants (Basel) 2023, 12(2).[PMID: 36829835]
8. Yoshida A et al: Recognition of G-quadruplex RNA by a crucial RNA methyltransferase component, METTL14. Nucleic Acids Res 2022, 50(1):449-457.[PMID: 34908152]
9. Jara-Espejo M, Fleming AM, Burrows CJ: Potential G-Quadruplex Forming Sequences and N(6)-Methyladenosine Colocalize at Human Pre-mRNA Intron Splice Sites. ACS Chem Biol 2020, 15(6):1292-1300.[PMID: 32396327]
10. Anastasakis DG et al: Nuclear PKM2 binds pre-mRNA at folded G-quadruplexes and reveals their gene regulatory role. Mol Cell 2024, 84(19):3775-3789 e3776.[PMID: 39153475]
11. Kharel P, Ivanov P: PKM2-G-quadruplex interactions conspire to regulate the cancer transcriptome. Mol Cell 2024, 84(19):3574-3575.[PMID: 39366344]
12. Matsuo K et al: RNA G-quadruplexes form scaffolds that promote neuropathological alpha-synuclein aggregation. Cell 2024, 187(24):6835-6848 e6820.[PMID: 39426376]
13. Dumas L et al: G-Quadruplexes in RNA Biology: Recent Advances and Future Directions. Trends Biochem Sci 2021, 46(4):270-283.[PMID: 33303320]
14. Kwok CK et al. rG4-seq reveals widespread formation of G-quadruplex structures in the human transcriptome. Nat. Methods 13, 841�C844 (2016).
15. Yang SY et al. Transcriptome-wide identification of transient RNA G-quadruplexes in human cells. Nat. Chem. Biol. 14, 180�C183 (2018).
16. Yeung PY et al. Systematic evaluation and optimization of the experimental steps in RNA G-quadruplex structure sequencing. Sci. Rep. 9, 8091 (2019).
17. Weng X et al. Keth-seq for transcriptome-wide RNA structure mapping. Nat. Chem. Biol. 16, 489�C492 (2020).
18. Hansel-Hertsch R et al. G-quadruplex structures mark human regulatory chromatin. Nat. Genet. 48, 1267�C1272 (2016).
19. Herviou P et al. hnRNP H/F drive RNA G-quadruplex-mediated translation linked to genomic instability and therapy resistance in glioblastoma. Nat. Commun. 11, 2661 (2020).
20. Simko EAJ et al. G-quadruplexes offer a conserved structural motif for NONO recruitment to NEAT1 architectural lncRNA. Nucleic Acids Res. 48, 7421�C7438 (2020).
21. Bolduc F et al. The small nuclear ribonucleoprotein polypeptide A (SNRPA) binds to the G-quadruplex of the BAG-1 5'UTR. Biochimie 176, 122�C127 (2020).
22. Guo JU et al. RNA G-quadruplexes are globally unfolded in eukaryotic cells and depleted in bacteria. Science. Sep 23;353(6306):aaf5371 (2016).
23. von Hacht A et al. Identification and characterization of RNA guanine-quadruplex binding proteins. Nucleic Acids Res. Jun;42(10):6630-44 (2014).
24. Haeusler et al. C9orf72 nucleotide repeat structures initiate molecular cascades of disease. Nature 507, 195�C200 (2014).
25. McRae EKS et al. Human DDX21 binds and unwinds RNA guanine quadruplexes. Nucleic Acids Res. Jun 20;45(11):6656-6668 (2017).
26. Serikawa T et al. Comprehensive identification of proteins binding to RNA G-quadruplex motifs in the 5' UTR of tumor-associated mRNAs. Biochimie. Jan;144:169-184 (2018).
27. Herdy B et al. Analysis of NRAS RNA G-quadruplex binding proteins reveals DDX3X as a novel interactor of cellular G-quadruplex containing transcripts. Nucleic Acids Res. Nov 30;46(21):11592-11604 (2018).
28. Yu H et al. G4Atlas: a comprehensive transcriptome-wide G-quadruplex database. Nucleic Acids Res. Jan 6;51(D1):D126-D134 (2023).
29. Bourdon S et al. QUADRatlas: the RNA G-quadruplex and RG4-binding proteins database. Nucleic Acids Res. Jan 6;51(D1):D240-D247 (2023).
30. Qian SH et al. EndoQuad: a comprehensive genome-wide experimentally validated endogenous G-quadruplex database. Nucleic Acids Res. Jan 5;52(D1):D72-D80 (2024).
31. Zhong HS et al. G4Bank: A database of experimentally identified DNA G-quadruplex sequences. Interdiscip Sci. Sep;15(3):515-523 (2023).
32. Birney E et al: An overview of Ensembl. Genome Res. May;14(5):925-8 (2004).
�پW(w��ng)��www.aksomics.com
�Ԓ��800-820-5058 400-886-5058 021-64451989
��ַ���Ϻ����h�Ѕ^(q��)��й�·2168̖(h��o)�ֽ��ǻۏV��(ch��ng)10C��4��




���Մ�(d��ng)�B(t��i) | �˲��Ј�(ch��ng) | �¼��g(sh��)���� | �Ї�(gu��)�ƌW(xu��)�� | ��չ�_(t��i) | BioHot | ���v��ֱ�� | ��(hu��)չ���� | �r(ji��)���� | ���g(sh��)��Ӎ | ���M(f��i)ԇ��
���(qu��n)���� ����ͨ
Copyright© eBiotrade.com, All Rights Reserved
(li��n)ϵ���䣺
��ICP��09063491̖(h��o)